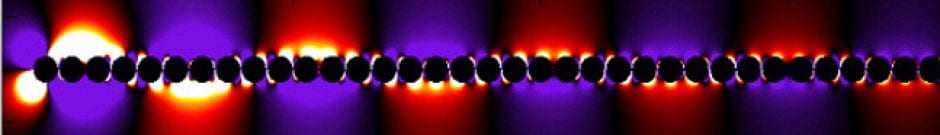
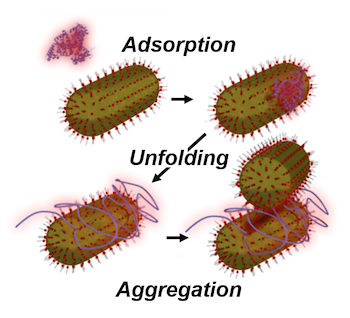
The fate of nanoparticles interacting with the human body is strongly influenced by a protein corona that forms when the nanoparticles come into contact with biological fluids. The type and number of proteins that absorb strongly depends on the particle size and surface chemistry and furthermore can change over the time period that the nanoparticles stay in the body. In situ, spectroscopic techniques, ideally at the single particle/molecule level, are therefore needed. The Link lab has developed correlation spectroscopy approaches utilizing the plasmon resonance to determine changes in the nanoparticle hydrodynamic radius due to protein binding, and together with collaborators (Prof. Landes, Rice) is using super-resolution fluorescence imaging to identify binding of individual dye-labeled proteins. To date, we have focused on bovine serum albumin (BSA) as it is the most abundant protein in blood. We have found that on negatively charged nanoparticles BSA forms a monolayer, which protects the nanoparticles from aggregation even at high salt concentration. However, at low BSA concentrations and for positively charged nanoparticles we surprisingly observed that BSA unfolds and then triggers nanoparticle aggregations. The ultimate goal of these studies is to understand the competitive binding of all serum proteins in real time while nanoparticles interact with cells for the safe use of nanomaterials and to potentially engineer protein coronas for enhanced medical applications.
Selected Publications:
- S. Dominguez-Medina, L. Kisley, L. J. Tauzin, A. Hoggard, B. Shuang, A. S. D. S. Indrasekara, S. Chen, L. -Y. Wang, P. J. Derry, A. Liopo, E. R. Zubarev, C. F. Landes, and S. Link Adsorption and Unfolding of a Single Protein Triggers Nanoparticle Aggregation. ACS Nano 10, 2103 (2016) link
- S. Dominguez-Medina, S. Chen, L. J. Blankenburg, P. Swanglap, C. F. Landes, and S. Link Measuring the Hydrodynamic Size of Nanoparticles using Fluctuation Correlation Spectroscopy. Annual Review of Physical Chemistry, 67, 489 (2016) link
Please also visit All Publications.